Publications, Posters & Presentations

The Almac claraT Total mRNA report has been utilised by multiple cancer researchers to enable their gene expression biomarker analysis. The following are some of the Publications, Posters and Presentations in which the researchers used claraT across various cancer indications.
Almac is delighted to collaborate with and support these research institutions and global biopharma companies to advance human health.
We thank them for their kind permission to include their work here.
Publications
University of Kansas Medical Center, USA
ASCO JCO Precision Oncology
“Dual Prognostic Classification of Triple-Negative Breast Cancer by DNA Damage Immune Response and Homologous Recombination Deficiency.”
Dr Shane R. Stecklein
The Study, led by Dr Stecklein and colleagues demonstrated that the classification of DDIR assay and HRD could be used as a dual prognostic classifier in triple negative breast cancer (TNBC) patients. In summary, immune active (DDIR+/HRD+) TNBCs exhibit the most favourable prognosis whereas immune depleted (DDIR-/HRD-) show the worst prognosis. There is also an intermediate prognosis sub-group whereby being positive for only one biomer (i.e. either DDIR or HRD) showed inferior prognosis to immune active TNBCs but superior prognosis to immune depleted TMBCs. Taken together these results could help refine the categorisation of TNBC patients of breast cancer patients.
View publication
Queens University Belfast
British Journal of Cancer
“Activation of a cGAS-STING-mediated immune response predicts response to neoadjuvant chemotherapy in early.” Dr. Stuart A. McIntosh
The Study led by Dr McIntosh and colleagues demonstrated that the DDIR assay could be used to identify a group of breast cancer patients which are more likely to benefit from a DNA damaging chemotherapy prior to surgery and have baseline activation of immune signalling. Furthermore, the team used a combination of transcriptome profiling, de novo discovery and the claraT report to demonstrate that patients with low baseline immune signalling or “cold” tumours are immune primed the through the use of this chemotherapy. Taken together these results could help refine the treatment of breast cancer patients and lead to better therapeutic outcomes.
View publication
University of Arkansas for Medical Sciences
MDPI Cancers
“Genomic and Transcriptomic Profiling of Brain Metastases.” Dr. Analiz Rodriguez
The Study led by Dr Rodriguez and colleagues utilised claraT as a method to facilitate the molecular profiling of brain metastases. Together with other genomic and transcriptomic tools, claraT helped identify novel molecular subtypes and identified potential new drivers of brain cancer progression in parallel to novel drug candidates that may promote improved responses in these patients.
View publication
Posters
Queens University Belfast- ASCO Symposium 2020
“Translational analysis of Esophageal Adenocarcinoma (EAC) patients treated with oxaliplatin and capecitabine (Xelox) +/-the dual Erb B inhibitor AZD8931 in the DEBIOC study.”
Dr Anita Lavery & Dr Richard Turkington
Download poster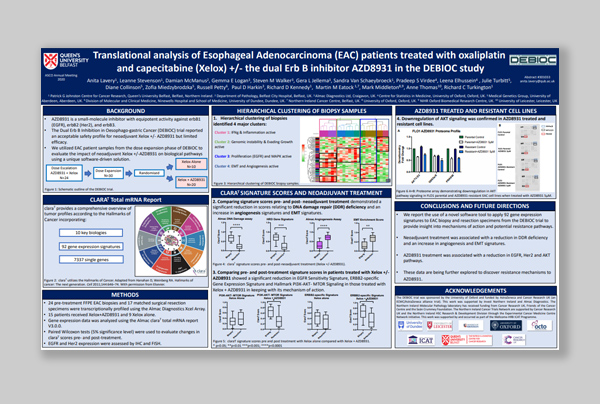
Almac Diagnostic Services
“Consensus Gene Expression Analysis to Identify Key Hallmarks of Cancer in Malignant Melanoma.”
Dr Jonathan Young
Download poster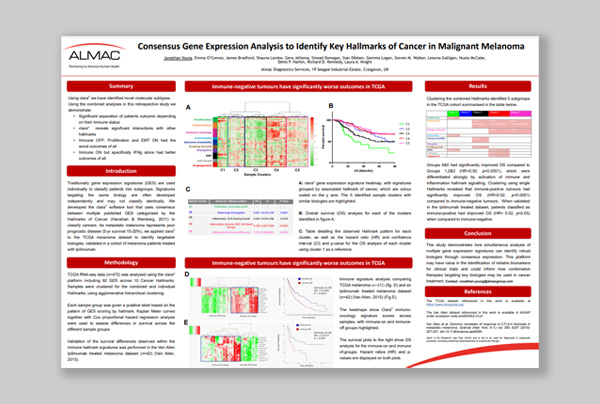
Gilead – ASCO Gastrointestinal Cancers Symposium 2020
“Identification of Cancer Hallmarks Associated with Benefit in Advanced Gastroesophageal Adenocarcinoma Patients Treated with Checkpoint Blockade.”
Emon Elboudwarej
Download poster
Presentations
SWOG – San Antonio Breast Cancer Symposium (SABCS) 2020
“Classification of triple negative breast cancer (TBNC) by DNA damage immune response (DDIR) signature and homologous recombination deficiency (HRD) status: Analysis of SWOG S9313 adjuvant trial.”
Shane Stecklein, University of Kansas
Download presentation
