


Whole Genome Methylation Sequencing (WGMS) Service



Whole Genome Methylation Sequencing (WGMS) Service
Whole Genome Methylation Sequencing (WGMS) Service
A high quality, robust Next Generation Sequencing (NGS) solution built on New England Biolab’s enzymatic methylation sequencing platform (EM-seq). Almac’s wet lab optimised workflow generates comprehensive WGMS data from fresh frozen (FF) samples. The platform covers coding and non-coding regions, generating raw sequencing data for use in biomarker discovery and retrospective clinical investigation. Optional bioinformatics services include processing of raw sequencing data to generate readily interpretable CpG site methylation profiles, and fully customised downstream analysis and reporting solutions.
For Research Use Only.
Robust Almac WGMS Data Generation Protocol:
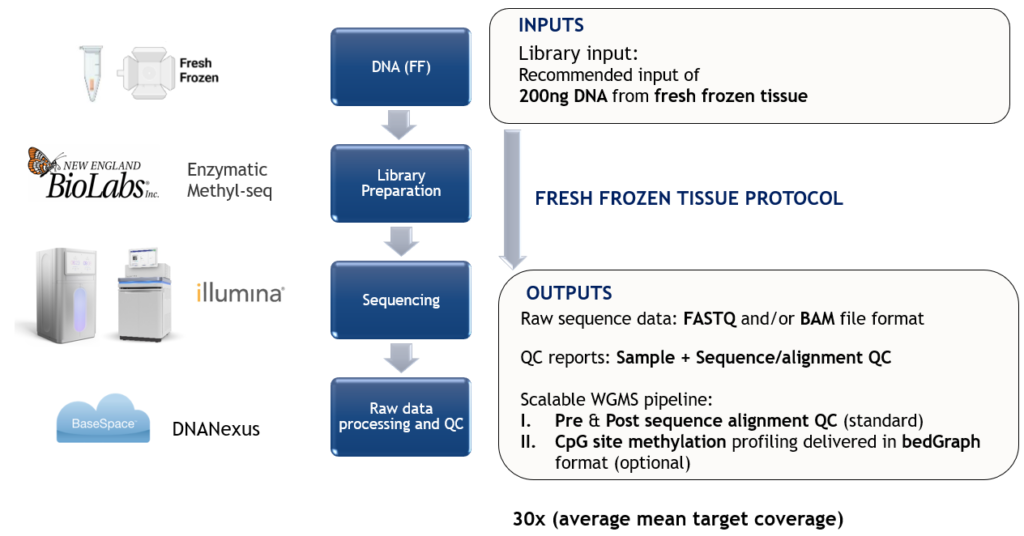
LAUNCH PROMOTIONAL OFFER – Initial customers adopting Almac WGMS service can avail of free readily interpretable CpG site methylation profiles for a limited time only.
Technical Specification and Performance Validation:
Almac analytical validation data confirms the highly accurate, repeatable and reproducible performance of the WGMS service.
For full technical specification & further validation information download the fact sheet.
Benefits:
- QC guarantee of 30X mean target coverage across sample cohort enables comprehensive downstream analysis and data interpretation.
- EM-Seq methodology allows for more uniform coverage of CpG site methylation sites.
- Protocol optimised for fresh frozen tissue samples.
- Interactive QC report enables rapid and comprehensive sample quality interrogation across the cohort.
- Raw Sequence data delivered in FASTQ format, compatible with the majority of bioinformatics pipelines for processing and interpreting Em-Seq data.
- Optional bioinformatics services to generate readily interpretable CpG site methylation profiles, and fully customised downstream analysis and reporting solutions
Contact us:
Interested in Almac’s Whole Genome Methylation Sequencing (WGMS) Service? Submit your details below:

